Algorithms Third Edition in C++ Part 5. Graph Algorithms (2006)
CHAPTER NINETEEN
Digraphs and DAGs
19.1 Glossary and Rules of the Game
Our definitions for digraphs are nearly identical to those in Chapter 17 for undirected graphs (as are some of the algorithms and programs that we use), but they are worth restating. The slight differences in the wording to account for edge directions imply structural properties that will be the focus of this chapter.
Definition 19.1 A digraph (or directed graph ) is a set of vertices plus a set of directed edges that connect ordered pairs of vertices (with no duplicate edges). We say that an edge goes from its first vertex to its second vertex.
As we did with undirected graphs, we disallow duplicate edges in this definition but reserve the option of allowing them when convenient for various applications and implementations. We explicitly allow self-loops in digraphs (and usually adopt the convention that every vertex has one) because they play a critical role in the basic algorithms.
Definition 19.2 A directed path in a digraph is a list of vertices in which there is a (directed) digraph edge connecting each vertex in the list to its successor in the list. We say that a vertex t is reachable from a vertex s if there is a directed path from s to t.
We adopt the convention that each vertex is reachable from itself and normally implement that assumption by ensuring that self-loops are present in our digraph representations.
Understanding many of the algorithms in this chapter requires understanding the connectivity properties of digraphs and the effect of these properties on the basic process of moving from one vertex to another along digraph edges. Developing such an understanding is more complicated for digraphs than for undirected graphs. For example, we might be able to tell at a glance whether a small undirected graph is connected or contains a cycle; these properties are not as easy to spot in digraphs, as indicated in the typical example illustrated in Figure 19.3.
Figure 19.3 A grid digraph
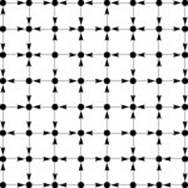

This small digraph is similar to the large grid network that we first considered in Chapter 1, except that it has a directed edge on every grid line, with the direction randomly chosen. Even though the graph has relatively few nodes, its connectivity properties are not readily apparent. Is there a directed path from the upper left corner to the lower right corner?
While examples like this highlight differences, it is important to note that what a human considers difficult may or may not be relevant to what a program considers difficult—for instance, writing a DFS class to find cycles in digraphs is no more difficult than for undirected graphs. More important, digraphs and graphs have essential structural differences. For example, the fact that t is reachable from s in a digraph indicates nothing about whether s is reachable from t. This distinction is obvious, but critical, as we shall see.
As mentioned in Section 17.3, the representations that we use for digraphs are essentially the same as those that we use for undirected graphs. Indeed, they are more straightforward because we represent each edge just once, as illustrated in Figure 19.4. In the adjacency-lists representation, an edge s-t is represented as a list node containing t in the linked list corresponding to s. In the adjacency-matrix representation, we need to maintain a full V -by-V matrix and to represent an edge s-t by a 1 bit in row s and column t. We do not put a 1 bit in row t and column s unless there is also an edge t-s. In general, the adjacency matrix for a digraph is not symmetric about the diagonal.
Figure 19.4 Digraph representations
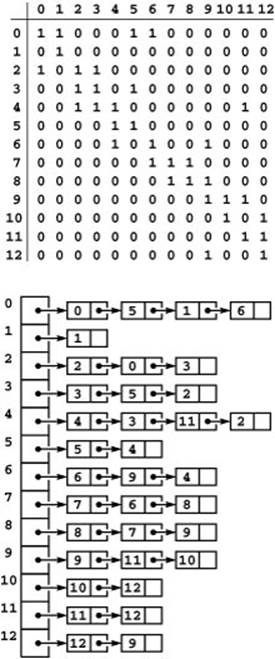
The adjacency-matrix and adjacency-lists representations of a digraph have only one representation of each edge, as illustrated in the adjacency-matrix (top) and adjacency-lists (bottom) representation of the graph depicted in Figure 19.1. These representations both include self-loops at every vertex, which is typical in digraph processing.
There is no difference in these representations between an undirected graph and a directed graph with self-loops at every vertex and two directed edges for each edge connecting distinct vertices in the undirected graph (one in each direction). Thus, we can use the algorithms that we develop in this chapter for digraphs to process undirected graphs, provided that we interpret the results appropriately. In addition, we use the programs that we considered in Chapter 17 as the basis for our digraph programs: Our DenseGRAPH and SparseMultiGRAPH class implementations Programs 17.7 through 17.10 build digraphs when the constructor has true as a second argument.
The indegree of a vertex in a digraph is the number of directed edges that lead to that vertex. The outdegree of a vertex in a digraph is the number of directed edges that emanate from that vertex. No vertex is reachable from a vertex of outdegree 0, which is called a sink; a vertex of indegree 0, which is called a source, is not reachable from any other vertex. A digraph where self-loops are allowed and every vertex has outdegree 1 is called a map (a function from the set of integers from 0 to V − 1 onto itself). We can easily compute the indegree and outdegree of each vertex, and find sources and sinks, in linear time and space proportional to V, using vertex-indexed vectors (see Exercise 19.19).
Program 19.1 Reversing a digraph
This function adds the edges of the digraph in its first argument to the digraph in its second argument, with their directions reversed. It uses two template parameters, so that the graphs can have different representations.
template <class inGraph, class outGraph>
void reverse(const inGraph &G, outGraph &R)
{
for (int v = 0; v < G.V(); v++)
{ typename inGraph::adjIterator A(G, v);
for (int w = A.beg(); !A.end(); w = A.nxt())
R.insert(Edge(w, v));
}
}
The reverse of a digraph is the digraph that we obtain by switching the direction of all the edges. Figure 19.5 shows the reverse and its representations for the digraph of Figure 19.1. We use the reverse in digraph algorithms when we need to know from where edges come because our standard representations tell us only where the edges go. For example, indegree and outdegree change roles when we reverse a digraph.
Figure 19.5 Digraph reversal
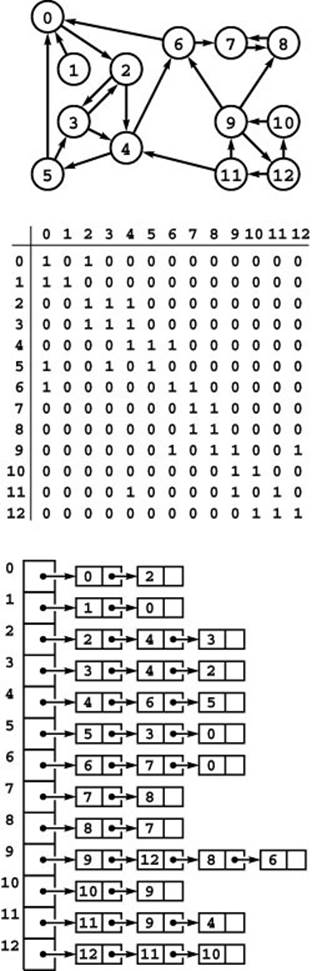
Reversing the edges of a digraph corresponds to transposing the adjacency matrix but requires rebuilding the adjacency lists (see Figures 19.1 and 19.4).
For an adjacency-matrix representation, we could compute the reverse by making a copy of the matrix and transposing it (interchanging its rows and columns). If we know that the graph is not going to be modified, we can actually use the reverse without any extra computation by simply interchanging the vertices in our references to edges when we want to refer to the reverse. For example, an edge s-t in a digraph G is indicated by a 1 in adj[s][t]. Thus, if we were to compute the reverse R of G, it would have a 1 in adj[t][s]. We do not need to do so, however, if we base our client implementations on the edge test edge(s, t), because to switch to the reverse we just replace every such reference by edge(t, s). This opportunity may seem obvious, but it is often overlooked. For an adjacency-lists representation, the reverse is a completely different data structure, and we need to take time proportional to the number of edges to build it, as shown in Program 19.1.
Yet another option, which we shall address in Chapter 22, is to maintain two representations of each edge, in the same manner as we do for undirected graphs (see Section 17.3) but with an extra bit that indicates edge direction. For example, to use this method in an adjacency-lists representation we would represent an edge s-t by a node for t on the adjacency list for s (with the direction bit set to indicate that to move from s to t is a forward traversal of the edge) and a node for s on the adjacency list for t (with the direction bit set to indicate that to move from t to s is a backward traversal of the edge). This representation supports algorithms that need to traverse edges in digraphs in both directions. It is also generally convenient to include pointers connecting the two representations of each edge in such cases. We defer considering this representation in detail to Chapter 22, where it plays an essential role.
In digraphs, by analogy to undirected graphs, we speak of directed cycles, which are directed paths from a vertex back to itself, and simple directed paths and cycles, where the vertices and edges are distinct. Note that s-t-s is a cycle of length 2 in a digraph but that cycles in undirected graphs must have at least three distinct vertices.
In many applications of digraphs, we do not expect to see any cycles, and we work with yet another type of combinatorial object.
Definition 19.3A directed acyclic graph (DAG) is a digraph with no directed cycles.
We expect DAGs, for example, in applications where we are using digraphs to model precedence relationships. DAGs not only arise naturally in these and other important applications, but also, as we shall see, in the study of the structure of general digraphs. A sample DAG is illustrated inFigure 19.6.
Figure 19.6 A directed acyclic graph (DAG)
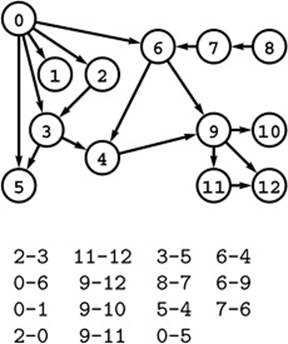
This digraph has no cycles, a property that is not immediately apparent from the edge list or even from examining its drawing.
Directed cycles are therefore the key to understanding connectivity in digraphs that are not DAGs. An undirected graph is connected if there is a path from every vertex to every other vertex; for digraphs, we modify the definition as follows:
Definition 19.4 A digraph is strongly connected if every vertex is reachable from every vertex.
The graph in Figure 19.1 is not strongly connected because, for example, there are no directed paths from vertices 9 through 12 to any of the other vertices in the graph.
As indicated by strongly, this definition implies a relationship between each pair of vertices stronger than reachability. In any digraph, we say that a pair of vertices s and t are strongly connected or mutually reachable if there is a directed path from s to t and a directed path from t to s. (Our convention that each vertex is reachable from itself implies that each vertex is strongly connected to itself.) A digraph is strongly connected if and only if all pairs of vertices are strongly connected. The defining property of strongly connected digraphs is one that we take for granted in connected undirected graphs: If there is a path from s to t, then there is a path from t to s. In the case of undirected graphs, we know this fact because the same path fits the bill, traversed in the other direction; in digraphs, it must be a different path.
Another way of saying that a pair of vertices is strongly connected is that they lie on some directed cyclic path. Recall that we use the term cyclic path instead of cycle to indicate that the path does not need to be simple. For example, in Figure 19.1, 5 and 6 are strongly connected because 6 is reachable from 5 via the directed path 5-4-2-0-6 and 5 is reachable from 6 via the directed path 6-4-3-5; and these paths imply that 5 and 6 lie on the directed cyclic path 5-4-2-0-6-4-3-5, but they do not lie on any (simple) directed cycle. Note that no DAG that contains more than one vertex is strongly connected.
Like simple connectivity in undirected graphs, this relation is transitive: If s is strongly connected to t, and t is strongly connected to u, then s is strongly connected to u. Strong connectivity is an equivalence relation that divides the vertices into equivalence classes containing vertices that are strongly connected to each other. (See Section 19.4 for a detailed discussion of equivalence relations.) Again, strong connectivity provides a property for digraphs that we take for granted with respect to connectivity in undirected graphs.
Figure 19.7 Digraph terminology
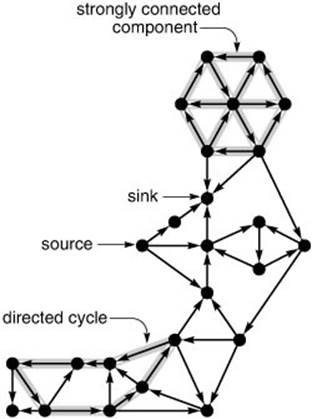
Sources (vertices with no edges coming in) and sinks (vertices with no edges going out) are easy to identify in digraph drawings like this one, but directed cycles and strongly connected components are more difficult to identify. What is the longest directed cycle in this digraph? How many strongly connected components with more than one vertex are there?
Property 19.1 A digraph that is not strongly connected comprises a set of strongly connected components ( strong components , for short), which are maximal strongly connected subgraphs, and a set of directed edges that go from one component to another.
Proof: Like components in undirected graphs, strong components in digraphs are induced subgraphs of subsets of vertices: Each vertex is in exactly one strong component. To prove this fact, we first note that every vertex belongs to at least one strong component, which contains (at least) the vertex itself. Then we note that every vertex belongs to at most one strong component: If a vertex were to belong to two different components, then there would be paths through that vertex connecting vertices in those components to each other, in both directions, which contradicts the maximality of both components.
For example, a digraph that consists of a single directed cycle has just one strong component. At the other extreme, each vertex in a DAG is a strong component, so each edge in a DAG goes from one component to another. In general, not all edges in a digraph are in the strong components. This situation is in contrast to the analogous situation for connected components in undirected graphs, where every vertex and every edge belongs to some connected component, but similar to the analogous situation for edge-connected components in undirected graphs. The strong components in a digraph are connected by edges that go from a vertex in one component to a vertex in another but do not go back again.
Property 19.2 Given a digraph D, define another digraph K ( D ) with one vertex corresponding to each strong component of D and one edge in K ( D ) corresponding to each edge in D that connects vertices in different strong components (connecting the vertices in K that correspond to the strong components that it connects in D). Then, K ( D ) is a DAG (which we call the kernel DAG of D).
Proof: If K ( D ) were to have a directed cycle, then vertices in two different strong components of D would fall on a directed cycle, and that would be a contradiction.
Figure 19.8 shows the strong components and the kernel DAG for a sample digraph. We look at algorithms for finding strong components and building kernel DAGs in Section 19.6.
Figure 19.8 Strong components and kernel DAG
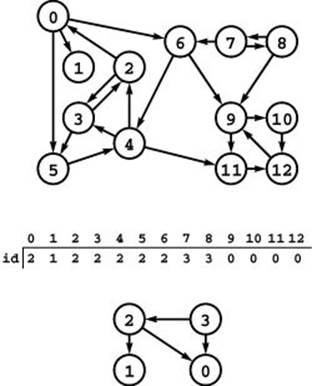
This digraph (top) consists of four strong components, as identified (with arbitrary integer labels) by the vertex-indexed array id (center). Component 0 consists of the vertices 9, 10, 11 and 12; component 1 consists of the single vertex 1; component 2 consists of the vertices 0, 2, 3, 4, 5, and 6; and component 3 consists of the vertices 7 and 8. If we draw the graph defined by the edges between different components, we get a DAG (bottom).
From these definitions, properties, and examples, it is clear that we need to be precise when referring to paths in digraphs. We need to consider at least the following three situations:
Connectivity We reserve the term connected for undirected graphs. In digraphs, we might say that two vertices are connected if they are connected in the undirected graph defined by ignoring edge directions, but we generally avoid such usage.
Reachability In digraphs, we say that vertex t is reachable from vertex s if there is a directed path from s to t. We generally avoid the term reachable when referring to undirected graphs, although we might consider it to be equivalent to connected because the idea of one vertex being reachable from another is intuitive in certain undirected graphs (for example, those that represent mazes).
Strong connectivity Two vertices in a digraph are strongly connected if they are mutually reachable; in undirected graphs, two connected vertices imply the existence of paths from each to the other. Strong connectivity in digraphs is similar in certain ways to edge connectivity in undirected graphs.
We wish to support digraph ADT operations that take two vertices s and t as arguments and allow us to test whether
• t is reachable from s
• s and t are strongly connected (mutually reachable)
What resource requirements are we willing to expend for these operations? As we saw in Section 17.5, DFS provides a simple solution for connectivity in undirected graphs that takes time proportional to V, but if we are willing to invest preprocessing time proportional to V + E and space proportional to V, we can answer connectivity queries in constant time. Later in this chapter, we examine algorithms for strong connectivity that have these same performance characteristics.
But our primary aim is to address the fact that reachability queries in digraphs are more difficult to handle than connectivity or strong connectivity queries. In this chapter, we examine classical algorithms that require preprocessing time proportional to V E and space proportional to V2, develop implementations that can achieve constant-time reachability queries with linear space and preprocessing time for some di-graphs, and study the difficulty of achieving this optimal performance for all digraphs.
Exercises
• 19.10 Give the adjacency-lists structure that is built by Program 17.9 for the digraph
3-7 1-4 7-8 0-5 5-2 3-8 2-9 0-6 4-9 2-6 6-4.
• 19.11 Write a program to generate random sparse digraphs for a well-chosen set of values of V and E such that you can use it to run meaningful empirical tests on digraphs drawn from the random-edges model.
• 19.12 Write a program to generate random sparse graphs for a well-chosen set of values of V and E such that you can use it to run meaningful empirical tests on graphs drawn from the random-graph model.
• 19.13 Write a program that generates random digraphs by connecting vertices arranged in a ![]() -by-
-by- ![]() grid to their neighbors, with edge directions randomly chosen (see Figure 19.3).
grid to their neighbors, with edge directions randomly chosen (see Figure 19.3).
• 19.14 Augment your program from Exercise 19.13 to add R extra random edges (all possible edges equally likely). For large R, shrink the grid so that the total number of edges remains about V. Test your program as described in Exercise 19.11.
• 19.15 Modify your program from Exercise 19.14 such that an extra edge goes from a vertex s to a vertex t with probability inversely proportional to the Euclidean distance between s and t.
• 19.16 Write a program that generates V random intervals in the unit interval, all of length d, then builds a digraph with an edge from interval s to interval t if and only if at least one of the endpoints of s falls within t (see Exercise 17.75). Determine how to set d so that the expected number of edges is E. Test your program as described in Exercise 19.11 (for low densities) and as described in Exercise 19.12 (for high densities).
• 19.17 Write a program that chooses V vertices and E edges from the real digraph that you found for Exercise 19.1. Test your program as described in Exercise 19.11 (for low densities) and as described in Exercise 19.12 (for high densities).
• 19.18 Write a program that produces each of the possible digraphs with V vertices and E edges with equal likelihood (see Exercise 17.70). Test your program as described in Exercise 19.11 (for low densities) and as described in Exercise 19.12 (for high densities).
• 19.19 Implement a class that provides clients with the capability to learn the indegree and outdegree of any given vertex in a digraph, in constant time, after linear-time preprocessing in the constructor. Then add member functions that return the number of sources and sinks, in constant time.
• 19.20 Use your program from Exercise 19.19 to find the average number of sources and sinks in various types of digraphs (see Exercises 19.11–18).
• 19.21 Show the adjacency-lists structure that is produced when you use Program 19.1 to find the reverse of the digraph
3-7 1-4 7-8 0-5 5-2 3-8 2-9 0-6 4-9 2-6 6-4.
• 19.22 Characterize the reverse of a map.
19.23 Design a digraph class that explicitly provides clients with the capability to refer to both a digraph and its reverse, and provide an implementation, for any representation that supports edge queries.
19.24 Provide an alternate implementation for your class in Exercise 19.23 that maintains both orientations of edges on adjacency lists.
• 19.25 Describe a family of strongly connected digraphs with V vertices and no (simple) directed cycles of length greater than 2.
• 19.26 Give the strong components and a kernel DAG of the digraph
3-7 1-4 7-8 0-5 5-2 3-8 2-9 0-6 4-9 2-6 6-4.
• 19.27 Give a kernel DAG of the grid digraph shown in Figure 19.3.
19.28 How many digraphs have V vertices, all of outdegree k?
• 19.29 What is the expected number of different adjacency-lists representations of a random digraph? Hint: Divide the total number of possible representations by the total number of digraphs.
All materials on the site are licensed Creative Commons Attribution-Sharealike 3.0 Unported CC BY-SA 3.0 & GNU Free Documentation License (GFDL)
If you are the copyright holder of any material contained on our site and intend to remove it, please contact our site administrator for approval.
© 2016-2025 All site design rights belong to S.Y.A.