Relevance Ranking for Vertical Search Engines, FIRST EDITION (2014)
Chapter 3. Medical Domain Search Ranking
Abstract
With the exponential growth of data stored in electronic health records (EHRs), it is imperative to identify effective means to help clinicians as well as administrators and researchers make full use of them. Recent research advances in natural language processing (NLP) have provided improved capabilities for automatically extracting concepts from narrative clinical documents. However, until these NLP-based tools become widely available and versatile enough to handle vaguely defined information retrieval needs of EHR users, a convenient and cost-effective solution continues to be in great demand. In this chapter, we introduce the concept of medical information retrieval, which provides medical professionals a handy tool to search among unstructured clinical narratives via an interface similar to that of general-purpose Web search engines, e.g., Google. In the latter part of the chapter, we also introduce several advanced features, such as intelligent, ontology-driven medical search query recommendation services and a collaborative search feature that encourages sharing of medical search knowledge among end users of EHR search tools.
Keywords
Electronic health record
EMERSE
Unified Medical Language system
search behavior analysis
relevance ranking
collaborative search
Introduction
Each patient visit to an outpatient care facility or hospital stay in an inpatient ward generates a great volume of data, ranging from physiological measurements to clinician judgments and decisions. Such data are not only important for reuse in current and later care episodes, they are also critical in supporting numerous secondary-use scenarios, including epidemic surveillance, population health management, and clinical and translational research [299,102]. It is widely believed that effective and comprehensive use of patient care data created in day-to-day clinical settings has the potential to transform the healthcare system into a self-learning vehicle to achieve better care quality, lower costs, and faster and greater scientific discoveries [157,115,128].
The launch of the health IT incentive program established through the American Recovery and Reinvestment Act of 2009 has stimulated widespread adoption of health IT systems in the United States, electronic health records (EHRs) in particular [37]. Medical professionals’ everyday interactions with such systems have in turn led to the exponential growth of rich, electronically captured data at the patient level, providing great promise for large-scale computational reuse. However, the increasing availability of electronic data does not automatically warrant the increasing availability of information [67]. A majority of clinical documents continue to exist in an unstructured, narrative format in the EHR era [260,352,52]; these documents are extremely difficult to process due to many characteristics unique to narrative medical data, such as frequent use of nonstandard terminologies and acronyms [340,365]. Recent studies have also shown that the quality of data stored in EHRs, compared to the quality of data recorded in paper forms, has deteriorated considerably due to the inappropriate use of electronic documentation features such as automated fill-in and copy-and-paste [160,149,369]. As a result, onerous, costly, and error-prone manual chart reviews are often needed in order to reuse the data in direct patient care or to prepare it for secondary-use purposes [159].
It is therefore imperative to identify effective means to help clinicians, as well as administrators and researchers, retrieve information from EHRs. Recent research advances in natural language processing (NLP) have provided improved capabilities for automatically extracting concepts from narrative clinical documents [411,353]. However, until these NLP-based tools become widely available and versatile enough to handle vaguely defined information retrieval needs of EHR users, a convenient and cost-effective solution continues to be in great demand. In this chapter, we introduce the concept of medical information retrieval, which provides medical professionals a handy tool to search among unstructured clinical narratives via an interface similar to that of general-purpose Web search engines such as Google. In the latter part of the chapter, we also introduce several advanced features, such as intelligent, ontology-driven medical search query recommendation services and a collaborative search feature that encourages sharing of medical search knowledge among end users of EHR search tools.
This chapter focuses on information retrieval systems for electronic health records in a clinical setting, to be distinguished from information retrieval systems for biomedical literature, such as PubMed, and those for consumer-oriented health information, such as MedlinePlus. Interested readers can refer to Hersh, 2009 [158], for information retrieval systems of biomedical literature and consumer-oriented information.
The chapter is organized as follows: In Section 3.1, we discuss the current research on EHR search engines and introduce a homegrown EHR search engine, EMERSE, which has been widely used at the University of Michigan Health Systems. Results of analyzing the search behaviors of EMERSE users are summarized in Section 3.2, which motivate the approaches to enhance the performance of EHR search. In Section 3.3, we describe effective approaches to enhance the relevance ranking of EHR search engines, which is a summary of the experiences of the TREC medical record track and the next generation of EMERSE. In Section 3.4, we present an alternative approach that leverages the collective intelligence instead of the machine intelligence to enhance medical record search.
3.1 Search Engines for Electronic Health Records
Clinicians and researchers routinely search medical records, but today they are doing so in a highly inefficient manner. Many of them simply go through each document manually and read through each clinical note to find the information they are looking for, a simple procedure known as chart review or chart abstraction. The tedious effort of manual search sometimes returns no results, either because the information was not there at all or because it was overlooked. An automatic search engine becomes essential in the modern setting of retrieving electronic health records.
Although there are multiple search engines to assist with searching medical literature or health-related Web pages [51,241,40,240,252,108,170,242], adoption of search engines for EHRs remains limited. This may, in part, be due to a lack of understanding of the information needs of clinicians and researchers compared to those of the general population. Additional factors limiting the widespread adoption of EHR search engines include the complex medical information contained in clinical documents and the inadequacy of standard search engines to meet users’ needs [270]. As a result, customized solutions are required. Even with such solutions, obtaining the proper approvals from a medical information technology department to integrate and support such a system and meeting all regulatory and privacy requirements are ongoing challenges that also limit the number of investigators who are able to work with such protected data in the clinical environment.
Only a few medical record search engines have been reported, and even among those it is difficult to know what level of adoption or usefulness has been achieved. The StarTracker system at Vanderbilt University was discussed in a brief report from 2003 [143]. At the time, it was available to 150 pilot users. It was reported to have been used successfully for cohort identification of clinical studies and to help with physician evaluation of clinical outcomes.
Columbia University also has a search engine, CISearch, that has been integrated with its locally developed EHR, WebCIS, since 2008 [263]. Supporting the notion that the information needs of an EHR search engine differ from those of standard search engines, the CISearch tool limits searches to a single patient at a time, and the system does not rank documents but rather displays the results in reverse chronological order, which is a common data view in medical record systems. The system was reported to have incorporated a limited number of document types, including discharge summaries, radiology reports, and pathology reports.
Other research systems have also been reported to support searching clinical documents, although these are not explicitly labeled as search engines [238]. An example is the STRIDE system at Stanford University, which has been shown to be of benefit for clinical decision making [126]. Additionally, the Informatics for Integrating Biology and the Bedside (i2b2) Workbench tool has been modified to handle searching free text notes [417].
At the University of Michigan we have had a search engine in our production environment since 2005. The Electronic Medical Record Search Engine (EMERSE) was developed to support users with an efficient and accurate means of querying our repository of clinical documents [145]. As of April 2013, we had over 60 million clinical documents and reports from approximately 2 million patients. EMERSE is used for a wide variety of information retrieval tasks, including research (e.g., cohort identification, eligibility determination, and data abstraction), quality improvement and quality assurance initiatives, risk management, and infection control monitoring.
From a technical perspective, EMERSE utilizes Apache Lucene to index and retrieve documents, but the Web application itself has been modified substantially to meet the needs of medical search. For example, stop words are not removed from the index, since many are themselves important acronyms. Examples include AND (axillary node dissection), ARE (active resistance exercise), and IS (incentive spirometry). Additionally, two indices are maintained of all documents: a standard lowercased index for the default case-insensitive searches and a case-sensitive index so users can distinguish medical terms from potential false positives. An example is the need to distinguish ALL (acute lymphoblastic leukemia) from the common word all.
EMERSE contains a large vocabulary of synonyms and related concepts that are presented to users to expand their search queries. For example, searching for “ibuprofen” would bring up suggestions that include brand names such as “Advil” and “Motrin” as well as common misspellings derived from our search logs, including “ibuprofin” and “ibuprophen.” As of April 2013, the synonym list contained 45,000 terms for about 11,000 concepts. From prior experimental work on intelligent query expansion, supported by the National Library of Medicine, we have learned that users often want fine-grained control over the terms used in their searches, since this can help them refine the search to meet their specific information needs. Incorporating other information sources from the Unified Medical Language System (UMLS), such as the [103], could further enhance users’ ability to expand their search queries. Assessing the relevance of documents at the concept level is a crucial practice in EHR information retrieval, which we will elaborate on in Section 3.3.
There are many aspects of clinical documents that make information retrieval challenging. These include the use of ambiguous terminology, known as hedge phrases (e.g., “the patient possibly has appendicitis”), as well as negation (e.g., “there is no evidence of appendicitis in this patient”). EMERSE provides a mechanism for handling negation that is easy for users to implement, but that deviates from standard practices for “typical” search engines. Negation can be achieved by adding exclude phrases, which are phrases that contain a concept of interest but in the wrong context. These phrases are notated with a minus sign in front of them, which tells the system to ignore those specific phrases but not the overall document itself. Thus, one can look for “complications” but ignore “no complications occurred.” Indeed, this was recently done for the Department of Ophthalmology as part of a quality assurance initiative to identify post-operative complications. The term “complication” or “complications” was searched and a collection of approximately 30 negated phrases were excluded. This greatly reduced the false-positive rate for the searches and allowed the department to conduct an efficient and accurate search for potential complications.
The EMERSE system also provides a framework to encourage the collaborative sharing and use of queries developed by others [405]. Saved searches, called bundles, can be created by any user and either shared or kept private. Many users have shared their bundles, and many bundles have been used by a wide variety of users. This paradigm allows users with specific medical expertise to share their knowledge with other users of the system, allowing those with less domain expertise to benefit from those with more. Bundles also provide a means for research teams to standardize their searches across a large group to ensure that the same processes are applied to each medical record [92]. We believe social search features an important functionality of the next generation of medical search engines. More details about the collaborative search component of EMERSE are provided in Section 3.4.
Studies have been carried out to assess various aspects of EMERSE. In addition to the collaborative search feature already described, we analyzed the EMERSE query logs to derive insights about users’ information-seeking habits [381]. Results of this query log analysis shed light on many important issues of medical search engines, which are elaborated in Section 3.2. Another study looked at how well eligibility determination for a psychiatric study could be carried out using traditional manual chart reviews in the EHR (which often requires clicking individually on each document to read through it) compared to using EMERSE [309]. Using EMERSE was far more efficient than the manual processes, yet the accuracy was maintained.
Another study looked into the feasibility of using a tool such as EMERSE to collect data for the American College of Surgeons, National Surgical Quality Improvement Program (ACS NSQIP) [146]. EMERSE was used to identify cases of post-operative myocardial infarction pulmonary embolus, a process that is traditionally performed manually. Utilizing the negation components of EMERSE was essential in these tasks to rule out false-positive results. Overall the system worked well, and it even identified cases that had been missed during the old standard manual review process. For myocardial infarction, sensitivity and specificity of 100.0% and 93.0%, respectively, were achieved. For pulmonary embolus, the sensitivity and specificity were 92.8% and 95.9%, respectively.
In the next section, we discuss the analysis of EHR search engine users’ search behaviors, especially users of EMERSE. The results of these analyses motivate various important approaches to enhancing the performance of EHR search engines.
3.2 Search Behavior Analysis
One might initially think that clinicians (physicians and nurses) would be the primary users of an EHR search engine. However, this is only partly true, according to the results of a survey of EMERSE users. EMERSE users cover nearly all medical disciplines, ranging from primary care fields such as general pediatrics, family medicine, and general internal medicine to subspecialties such as neurosurgery, urology, and gastroenterology. Among the users who responded to an EMERSE survey, only 7% are full-time practicing clinicians; the others are cancer registrars, infection control specialists, quality assurance officers, pharmacists, data managers, and other research-oriented individuals, including medical fellows, residents, students, and even undergraduates working on research projects within the University of Michigan Health System.
According to the survey, EMERSE users were using the search engine for various kinds of tasks. Two-thirds used the search engine to determine medication use for patients. Nearly as many users reported using EMERSE for assisting with clinical trials. Other uses included detection of adverse events, determining eligibility for clinical studies, infection surveillance, internal quality assurance projects, study feasibility determination, billing/claims abstraction, and risk management review. As a comparison, researches of the Columbia University’s EHR search engine revealed that searches for laboratory or test results and disease or syndromes constituted the majority of the search purposes.
The availability of longitudinal collection of search logs of EHR search engines made it possible to quantitatively analyze EMERSE users’ search behaviors. Natarajan et al. analyzed the query log of an EHR search engine with 2,207 queries [263]. They found that among these queries, 14.5% were navigational queries and 85.1% were informational. Searching for laboratory results and specific diseases were users’ predominant information needs. In 2011, Yang et al. analyzed a much larger collection that consisted of 202,905 queries and 35,928 user sessions recorded by the EMERSE search engine over four years [381]. The collection represented the information-seeking behavior of 533 medical professionals. Descriptive statistics of the queries, a categorization of information needs, and temporal patterns of the users’ information-seeking behavior were reported [381]. The results suggest that information needs in medical searches are substantially more complicated than those in general Web searches.
Specifically, the frequency of medical search queries does not follow a power-law distribution, as that of Web search queries does. A medical search query contains five terms on average, which is two times longer than the average length of a Web search query. Users of the EHR search engine typically start with a very short query (1.7 terms on average) and end up with a much longer query through a search session. A session of EHR search is considerably longer than a session of Web search, in terms of both the time period (14.8 minutes on average) or the number of queries (5.64 on average). All of these points suggest that it is substantially more difficult for users to compose an effective medical search query than a general Web search query.
In what aspects are the medical search queries more difficult? It is reported that more than 30% of the query terms are not covered by a common English dictionary, a medical dictionary, or a professional medical ontology, compared to less than 15% of terms in Web searches. Furthermore, 2,020 acronyms appeared 55,402 times in the EMERSE query log, where 18.9% of the queries contain at least one acronym [381].
The low coverage of query terms by medical dictionaries not only implies the substantial difficulty of developing effective spell-check modules for EHR search, but it also suggests the need to seek beyond the use of medical ontologies to enhance medical information retrieval. One possible direction leads towards deeper corpus mining and natural language processing of electronic health records.
Moreover, the categorization of medical search queries is substantially different from those in Web searches. The well-established categorization of information needs in Web searches into navigational, informational, and transactional queries does not apply to medical search [45]. This calls for a new categorization framework for medical search queries. Yang et al. suggests one possible categorization that discriminates high-precision information needs from high-recall information needs [381]. In that work, a clinical ontology is used to categorize the semantics of queries into a few high-level medical concepts (see Table 3.1). According to their results, around 28% of the queries are concerned with clinical findings, which include subcategories such as diseases and drug reactions. Furthermore, 12.2% of the queries are concerned with pharmaceutical or biological products, followed by procedures (11.7%) and situation with explicit context (8.9%). In addition 10.4% of the EMERSE queries cannot be described by any of the categories. It is an interesting question to ask to what extent such a semantic categorization is useful in enhancing the performance of an EHR search engine.
Table 3.1
Distribution of categories of medical concepts in EMERSE queries.*
|
Category |
Coverage (%) |
|
Clinical finding |
28.0 |
|
Pharmaceutical/biological products |
12.2 |
|
Procedures |
11.7 |
|
Unknown |
10.4 |
|
Situation with explicit context |
8.9 |
|
Body structure |
6.5 |
|
Observable entity |
4.3 |
|
Qualifier value |
3.7 |
|
Staging and scales |
3.2 |
|
Specimen |
2.2 |
|
Physical object |
2.2 |
|
Event |
1.7 |
|
Substance |
1.5 |
|
Linkage concept |
1.0 |
|
Physical force |
0.6 |
|
Environment or geographical location |
0.6 |
|
Social context |
0.5 |
|
Record artifact |
0.4 |
|
Organism |
0.3 |
|
Special concept |
0.03 |
* Queries submitted to EMERSE are categorized into the top 19 medical concepts in the ontology structure of SNOMED-CT [327], a standard clinical terminology.
All the findings from the analysis of real users suggest that medical search queries are in general much more sophisticated than Web search queries. This difficulty imposes considerable challenges for users to accurately formulate their search strategies. There is an urgent need to design effective mechanisms to assist users in formulating and improving their queries. These findings motivate many important directions for improving EHR search engines. Now let’s discuss three particular directions: relevance ranking, query recommendation, and collaborative search.
3.3 Relevance Ranking
Conventional search engines for medical records (e.g., EMERSE) do not provide the functionality of ranking matched results as is commonly done by Web search engines. This largely contributes to the uniqueness of the information needs in medical search. Indeed, when users search for medical records, the concept of a “hit” is different where there are usually no “best document” or “most relevant” records to rank and display. Instead, the final “answer” depends heavily on the initial question and is almost always determined after the user views a collection of documents for a patient, not a single document.
The heavy dependence on manual reviewing is largely due to the uncertainty and ambiguity that is inherent in medical records. Consider an example: A clinician is trying to identify patients diagnosed with asthma. The diagnosis of asthma for a young child can be difficult. When a child with difficulty breathing is on an initial clinic visit, the clinical note may mention terms such as wheezing, coughing, reactive airway disease, and, perhaps, asthma. However, this does not mean that the patient actually has asthma, because many children may develop wheezing during an upper respiratory viral infection. Observations of documented recurrent episodes have to be done before one might conclude that a child truly has asthma, and one should also take into account the prescribed medications and the changes in the patient’s condition based on the use of those medications. Therefore, a single medical record mentioning asthma, regardless of how frequently it is mentioned in the document, cannot truly provide a confident diagnosis. Therefore, in conventional EHR searches, the requirement of a high recall is usually more important than a high precision. The ranking of search results is not critical, since all the retrieved records will be manually reviewed by the users.
It was not until recently that designers of EHR search engines started to adopt relevance ranking in the retrieval architecture. On one hand, the volume of electronic records is growing rapidly, making it difficult for users to manually review all the documents retrieved. On the other hand, the recent development of natural language processing for EHRs has made it possible to handle negation [261], uncertainty [147], and ambiguity [63] to a certain degree. Relevance ranking has become appealing in many scenarios of medical record search, especially in scenarios with a need for high precision, such as identifying patients for clinical trials. In the next section we examine the effort of a medical record search challenge organized by the Text REtrieval Conference (TREC), followed by an implementation we adopt for the next generation of the EMERSE search engine.
3.3.1 Insights from the TREC Medical Record Track
The Text REtrieval Conference (TREC), organized by the U.S. National Institute of Standards and Technology (NIST) every year since 1992, provides a standard testbed to evaluate particular information retrieval (IR) tasks. In 2011 and 2012, TREC organized a new track called the Medical Record Track that provides an IR challenge task to identify cohorts of patients fitting certain criteria from electronic health records, which are similar to those specified for participation in clinical studies [359].
The task in both years used a collection of electronic health records from the University of Pittsburgh NLP repository, which consisted of 93,551 de-identified clinical reports collected from multiple hospitals, including radiology reports, history and physicals, consultation reports, emergency department reports, progress notes, discharge summaries, operative reports, surgical pathology reports, and cardiology reports [359]. These clinical reports were aggregated into 17,264 visits, each of which corresponds to the single stay at a hospital of an individual patient. Query topics (35 in 2011 and 50 in 2012) of the track represent different case definitions, which vary widely in terms of linguistic complexity. Both the selection of the query topics and the relevance assessments were carried out by physicians who were also students in the Oregon Health & Science University (OHSU) Biomedical Informatics Graduate Program.
It is reported that the majority of the participating teams employed some sort of domain-specific vocabulary normalization, concept extraction, and/or query expansion [359]. This largely attributes to the informal and ambiguous language used in health records. Indeed, a given medical entity is usually referred to by a wide variety of acronyms, abbreviations, and informal designations [359]. A typical method used by the participating teams is to identify and normalize medical terms in text and map the terms to concepts in the UMLS metathesaurus using MetaMap. Some participants also map these UMLS concepts to entities in some controlled medical vocabularies, such as SNOMED-CT or MeSH, which allows the system to navigate through different granularities of concepts in the queries and documents and to expand the queries using related concepts in the ontology.
It is also worth mentioning that many other information sources were used for query expansion. These sources included domain specific taxonomy such as the International Classification of Diseases (ICD) codes, term relations mined from the EHR corpus, and various external sources such as encyclopedias, biomedical literature, and Wikipedia, as well as the open Web.
It is also important to note that medical records frequently use negated expressions to document the absence of symptoms, behaviors, and abnormal diagnostic results [359]. Results from the participating teams suggest that it is an effective practice to specifically handle negations in EHR retrieval, which is in contrast to many other ad hoc information retrieval tasks.
A team from Cengage Learning achieved top performance in the TREC 2011 medical track [202]. The key techniques they applied included information extraction and query expansion. Specifically, medical terms that appeared in the UMLS were extracted from the queries and the documents, together with a limited collection of demographical information. Three methods of query expansion were explored, one through UMLS-related terms, one through a network of terms built from UMLS, and the other through terms from in-house medical reference encyclopedias. It is worth mentioning that they conducted a specific treatment in information extraction to detect negations and uncertainty. Furthermore, 256 and 130 rules were used to detect negation uncertainty, respectively. Medical terms associated with a negated or uncertain expression were excluded from the index. Such a treatment was reported to have improved the average precision by 5%.
In the TREC 2012 medical track, a team from the University of Delaware achieved the top performance [414]. It is reported that they mainly explored three effective directions: (1) aggregating evidence of relevance at different granularities, i.e., at the level of individual medical reports and at the level of patient visits; (2) query expansion through a variety of large-scale external sources; and (3) applying machine learning methods to combine classical IR models and different types of features. A Markov random field (MRF) was employed to model the proximity dependency of terms.
To summarize, it is a common practice to enhance relevance ranking in medical records searches by introducing concept-level relevance assessment and query expansion. In the next subsection, we discuss our practice of the two directions in the next generation of the EMERSE search engine.
3.3.2 Implementing and Evaluating Relevance Ranking in EHR Search Engines
We built a proof-of-concept prototype of the next generation of EHR search engine (referred to as EHR-SE), which features a concept-level relevance ranking and a component of query recommendation. As a proof of concept, we adopted straightforward query recommendation and relevance ranking algorithms. Under the same architecture, we could apply more sophisticated query suggestion methods and/or relevance ranking methods.
In general, the new EHR search engine advances EMERSE and other existing full-text search engines for medical records by assessing document relevance and recommending alternative query terms at the level of medical concepts instead of at the level of individual words. A document is retrieved because one of the medical concepts implied by its terms matches the concepts in the query. Relevant documents are ranked based on how well the concepts in the documents match the concepts in the query, through classical retrieval methods extended to the concept level. For example, when the query is looking for patients with “hearing loss,” documents containing “difficulty of hearing” or “deaf” will also be retrieved and ranked highly.
As a proof of concept, we adopted the same corpus of electronic health records used in the TREC medical record track. Medical terms and concepts were extracted from the reports in two ways. First, medical terms were identified and mapped to the metathesaurus concepts in the UMLS by MetaMap, with a numerical score from 0 to 1000 representing the confidence in the match. Second, we also extracted medical terms that were frequently searched by EMERSE users and mapped them to an empirical synonym set (ESS), which was constructed through mining the search log of EMERSE. The medical records were then indexed by either the medical terms in the metathesaurus or the synonyms in the EES using the Lemur toolkit, denoted as indexes ![]() and
and ![]() , respectively.
, respectively.
With the new representation of an individual document or a query as a set of medical terms, the relevance score of a document, given a query, can be calculated based on a classical vector space retrieval model known as the method:
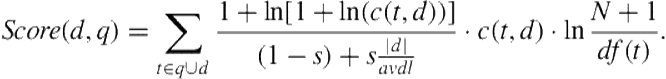 (3.1)
(3.1)
In Equation 3.1, ![]() , and
, and ![]() denote a medical term, a medical document, and a query, respectively;
denote a medical term, a medical document, and a query, respectively; ![]() denotes the frequency of the term
denotes the frequency of the term ![]() appearing in the document;
appearing in the document; ![]() denotes the length of the document normalized by the average length of documents in the collection, and
denotes the length of the document normalized by the average length of documents in the collection, and ![]() denotes the document frequency of
denotes the document frequency of ![]() (i.e., the number of documents containing the term
(i.e., the number of documents containing the term ![]() ). We modified the formula such that (1) the set of synonyms under the same concept are treated as a single term; (2) the metrics are calculated from both of the indexes (the metathesaurus index
). We modified the formula such that (1) the set of synonyms under the same concept are treated as a single term; (2) the metrics are calculated from both of the indexes (the metathesaurus index ![]() and the ESS index
and the ESS index ![]() ) and then combined; and (3) the metrics consider the case that a term may be contained in multiple metathesaurus concepts of ESS subsets.
) and then combined; and (3) the metrics consider the case that a term may be contained in multiple metathesaurus concepts of ESS subsets.
The analysis of the query log of EMERSE showed that the performance of medical record searches still suffers from the low quality of queries entered by end users and the tremendous amount of redundant effort on exploring similar queries across users [4]. It is difficult even for clinicians with years of training to form adequate queries that lead to desirable search results due to the complexity of the domain and the information needs [5]. Two techniques that have been deployed in commercial search engines could be helpful to this issue in the medical context: (1) query recommendations, which provide alternative query terms based on the “wisdom” of data mining algorithms; and (2) social searches, which select and promote efficient queries among users based on the “wisdom of the crowd.”
The new EHR-SE is featured with a component of recommending alternative query concepts to users. When a user submits a query ![]() , the same process for document preprocessing is applied to identify the medical terms in the query and match them to concepts. For each medical term
, the same process for document preprocessing is applied to identify the medical terms in the query and match them to concepts. For each medical term ![]() extracted from the query, the query recommendation algorithm simply generates all synonyms of the term that are either under the same metathesaurus concepts,
extracted from the query, the query recommendation algorithm simply generates all synonyms of the term that are either under the same metathesaurus concepts, ![]() , or belong to the same synonym subset in ESS,
, or belong to the same synonym subset in ESS, ![]() . The union of
. The union of ![]() and
and ![]() is presented to expand the original query. A particular example of the query expansion is shown in Figure 3.1, a snapshot of the real EHR-SE system.
is presented to expand the original query. A particular example of the query expansion is shown in Figure 3.1, a snapshot of the real EHR-SE system.
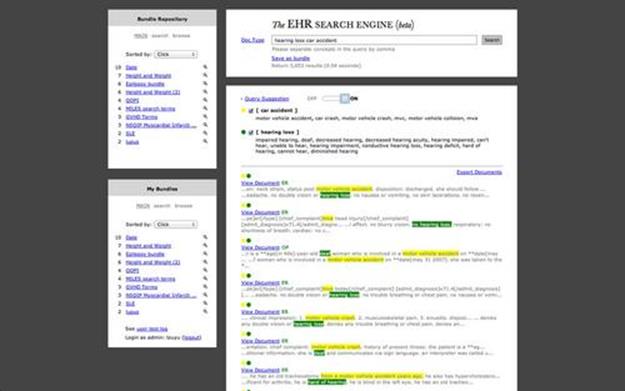
FIGURE 3.1 Snapshot of the new EHR Search Engine (EHR-SE). Medical terms in the query are expanded with synonyms under the same medical concepts. A user can choose to turn on or turn off the query suggestions. The retrieved results are ranked in the lower-right panel with the matched concepts highlighted. The left panel provides a functionality of social search so that the users can store, share, and adopt effective queries.
The effectiveness of this proof-of-concept system was evaluated with a carefully designed user study. 33 participants were recruited, representing different departments and divisions in the University of Michigan Health System.
We assigned standardized scenarios to each participant to simulate the real information needs in medical record search.
In each scenario, a participant could explore the system with as many queries as necessary. After obtaining satisfactory search results, each participant answered three questions with a score from 1 to 5 and provided narratives to describe their perceptions of the system performance.
Overall, the participants gave very positive feedback on the system performance. The overall evaluation score suggested that the performance of the system when automatic query recommendation was turned on was significantly better than the performance when the service was turned off.
In summary, concept-level relevance assessment and automatic query expansion play an important role in relevance ranking of medical record searches. In this way, the challenge introduced by the sophistication of the medical domain and the information needs are alleviated to a certain degree. However, there are still quite a few barriers to solving all the problems with automatic query expansion or recommendation. For example, findings from the TREC medical record track suggest that automatic query expansion for some query types may cause query drifts, which hurt retrieval performance [359]. In a recent paper summarizing the failure analysis of the TREC medical record track, the authors listed a few common causes of retrieval errors, including irrelevant chart references to medical terms; variation of the uses and spellings of terminology; ambiguity of different conditions with similar names; ambiguity among past, present, and future conditions or procedures, and imperfect treatments of negations and uncertainties [112].
In the long run, most of these barriers may be overcome with more advanced natural language processing techniques for electronic health records. An alternative approach to helping users formulate the effective queries, however, may go beyond machine intelligence and require the utilization of social intelligence. The following section describes our exploration of collaborative searches, which allows users to disseminate search knowledge in a social setting.
3.4 Collaborative Search
An intelligent search engine does not solve all problems of information retrieval for medical records; its performance depends highly on the quality of search queries that users are able to construct. From our analysis of the EMERSE query log, it seems that average users often do not have adequate knowledge to construct effective and inclusive search queries, given that users usually revise their queries multiple times through a search session and frequently adopt system-generated or socially disseminated suggestions.
One way to address this issue that has drawn increasing attention is the concept of social information seeking or collaborative search. Such a process enables users to collaboratively refine the quality of search queries as well as the quality of information resources. Examples include the Yahoo! Search Pad,1 Microsoft SearchTogether,2 and various social bookmarking sites. A common goal of these approaches is to encourage users to record the queries they found effective and share such search knowledge with other users.
Can this concept be applied to facilitate information retrieval for medical records? We implemented a “collaborative search” feature in EMERSE that allows users to preserve their search knowledge and share it with other users of the search engine. Through this feature, a user can create, modify, and share so-called “” that contain collections of keywords and regular expressions to describe that user’s information needs [8]. For example, a commonly used search term bundle, named “Cancer Staging Terms,” contains as many as 202 distinct search terms, such as “gleason,” “staging workup,” and “Tmic” [405]. Analysis of the EMERSE query log shows that as many as 27.7% of the search sessions ended up adopting a search bundle created by other users.
There are two ways for an EMERSE user to share a search bundle that user created. She may share it to the public, so that all EMERSE users can view and adopt the bundle. She may also share it exclusively to a list of users who can benefit from the bundle, according to her own assessment. Our survey found that 34.3% of all bundles were shared privately, and 20.5% of the bundles were shared publicly, with the rest not shared at all. About 41.9% of the EMERSE users were involved in collaborative search activities (either shared or adopted a bundle) based on either privately shared bundles (24.8%) or publicly shared bundles (26.8%).
To better describe the effect of the collaborative search feature, a social network analysis was conducted that quantified the utility of search bundles in enhancing the diffusion of search knowledge [405].
Figure 3.2 presents a network of search bundles created and adopted by users associated with different departments in the University of Michigan Health System. Clearly, the interdepartmental bundle-sharing activities (red links) played important roles in disseminating the search knowledge from one department to another. The potential of this feature was far from being fully realized, however, given that a larger number of bundles were still used only internally in a department [405].
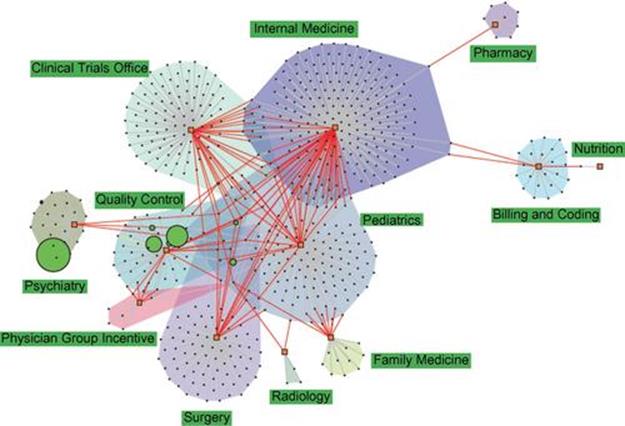
FIGURE 3.2 Network of search bundles presenting the diffusion of search knowledge across departments. Every circle node is a search bundle, sized proportionally to its frequency of use. Every rectangle node refers to a department, with colored convex hulls encompassing all search bundles created by the same department. Every red edge connects a bundle to the departments of the users who “borrowed” them in search; every gray edge connects a bundle to the department of the user who created it. For interpretation of the references to color in this figure legend, the reader is referred to the web version of this book.
Figure 3.3 presents a different view of the diffusion of search knowledge by including the users in the network.
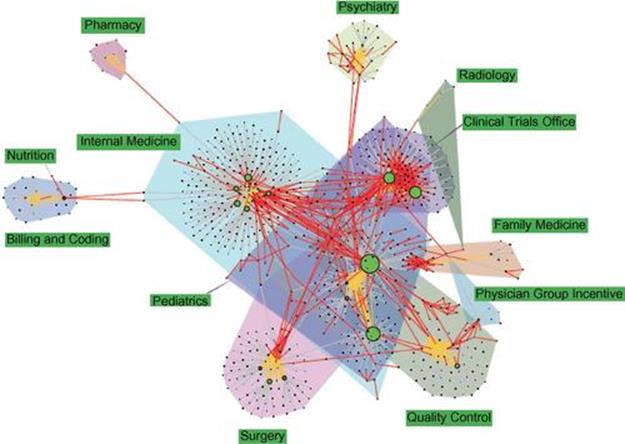
FIGURE 3.3 Network of users and bundles presenting the diffusion of search knowledge. Every circle node is a user, sized proportionally to the number of search bundles he has created. Orange edges connect users in the same department. Every other (rectangle) node is a search bundle. A colored convex hull encompasses all users and search bundles created by the users in the same department. Every red edge connects a bundle to the users who “borrowed” them in search; every gray edge connects a bundle to the user who created it.
To quantify and compare the effectiveness of publicly shared bundles and privately shared bundles in the diffusion of search knowledge, analysis was conducted on a few creator-consumer networks, the nodes of which represent EMERSE users, and a (directed) edge connects the creator of a bundle to the consumer of the bundle [405]. Overall, a creator-consumer network established through collaborative search presents a high average degree, a high clustering coefficient, and a small average shortest path compared to random networks of the same size. This indicates that the collaborative search feature has successfully facilitated the diffusion of search knowledge, forming a small-world network of search users. Between privately shared bundles and publicly shared bundles, the latter seems to be slightly more effective, with the combination of the two types of bundles significantly more effective than either individual type.
Our work also compared the search knowledge diffusion networks with a hypothetical social network of users constructed by connecting users who had ever typed in the same queries [405]. Such an analysis revealed a big potential gain if we had a better mechanism of collaborative search. The hypothetical network was featured with much fewer singletons, a much higher average degree, and a much shorter average shortest path length. The findings of this study suggest that although the collaborative search feature had effectively improved search-knowledge diffusion in medical record search, its potential was far from being fully realized.
3.5 Conclusion
Information retrieval of medical records in a clinical setting is especially challenging in various ways. The sophistication of the domain knowledge, the unique linguistic properties of the medical records, the complex information needs, and the barriers due to patient privacy protection have together limited the effectiveness of search engines for electronic health records. In this chapter we presented recent findings from the experience with our in-house EHR search engines and the TREC medical record track, which shed light on several effective practices for improving medical records search. Specifically, we highlighted three key approaches: concept-level relevance ranking, automatic query expansion, and collaborative search.
There remain many challenging issues in effective information retrieval for medical records. A more accurate and robust natural language processing pipeline for medical records should largely improve the performance of EHR search engines. Such a pipeline should provide powerful tools for medical concept extraction, acronym resolution, word disambiguation, and negation and uncertainty detection.
An even more urgent task is to better understand the information needs and search behaviors of the users of EHR search engines. In analogy to the query log analysis in Web search, which has long been proven a successful approach to enhancing the performance of Web search engines, better models and techniques utilizing the observations of search behaviors are anticipated to significantly enhance the effectiveness of EHR search engines.
Finally, this chapter pointed out a potentially high-return approach, collaborative search, that is an alternative to the methods based on machine intelligence. With such an approach, the users of medical record search engines are enabled to preserve, disseminate, and promote search knowledge that leads to satisfactory search results. A promising future direction of medical information retrieval will be incentive-centered designs that better engage users with the collaborative search features.
1 www.ysearchblog.com/2009/07/07/unveiling-yahoo-search-pad/![]() .
.
2 http://research.microsoft.com/en-us/um/redmond/projects/searchtogether/![]() .
.
All materials on the site are licensed Creative Commons Attribution-Sharealike 3.0 Unported CC BY-SA 3.0 & GNU Free Documentation License (GFDL)
If you are the copyright holder of any material contained on our site and intend to remove it, please contact our site administrator for approval.
© 2016-2026 All site design rights belong to S.Y.A.